ImmunoMap: A Novel Bioinformatics Tool for Immune Cell Repertoire Analysis

Abstract
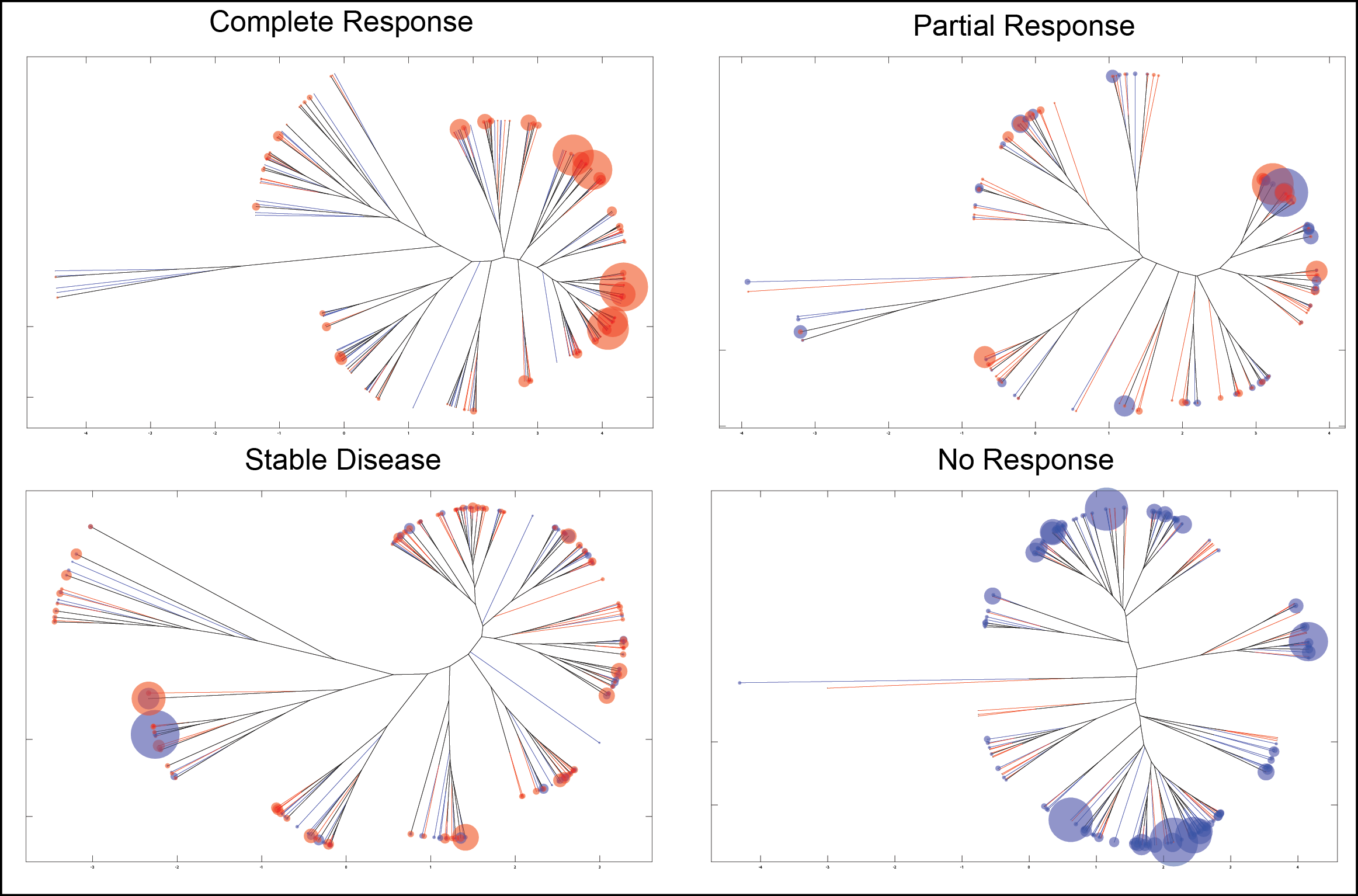
Poster Background: There has been a dramatic increase in T-cell Receptor (TCR) sequencing spurred, in part, by the widespread adoption of this technology across academic medical centers and by the rapid commercialization of TCR sequencing. While the raw TCR sequencing data has increased, there has been little in the way of approaches to parse the data in a biologically meaningful fashion. The ability to parse this new type of ‘big data’ quickly and efficiently to understand the T-cell repertoire in a structurally relevant manner has the potential to open the way to new discoveries about how the immune system is able to respond to insults such as cancer and infectious diseases. Materials and Methods: Here we describe a novel method utilizing phylogenetic and sequencing analysis to visualize and quantify TCR repertoire diversity (Figure 1). To demonstrate the utility of the approach, we have applied it to understanding the shaping of the CD8 T Cell response to self (TRP2) and foreign (SIY) antigens in naïve and tumor bearing B6 mice. Additionally, this method was applied to tumor infiltrating lymphocytes (TIL’s) from patients undergoing Nivolumab (anti-PD1) therapy in a BMS-038 clinical trial for metastatic melanoma to understand TCR repertoire characteristics between responders and non-responders. Results: Analysis of the naïve CD8 response to SIY showed a lower clonality yet more closely structurally related response whereas CD8 responses to TRP2 were highly clonal yet less structurally related. Presence of tumor demonstrated more immune pressure and selection on TRP2 with structurally different responses arising in the presence of tumor. In contrast, this type of immune pressure was not as pronounced in the SIY response. Finally within tumor-bearing animals, SIY response became more clonal yet remained homologous as the repertoire was analyzed closer to tumor site whereas TRP2 response became less clonal and less structurally related. In the BMS-038 trial data, this analysis revealed that patients whose CD8 response had a larger contribution from newly expanding TCR structures responded better to therapy. Conclusions: In summary, we have developed and demonstrated a novel method to meaningfully parse and interpret TCR repertoire data and have applied it to yield a novel understanding of CD8 T Cell responses to different types of antigens as well as key characteristics in those who respond to anti-PD1 therapy.
Figure 1
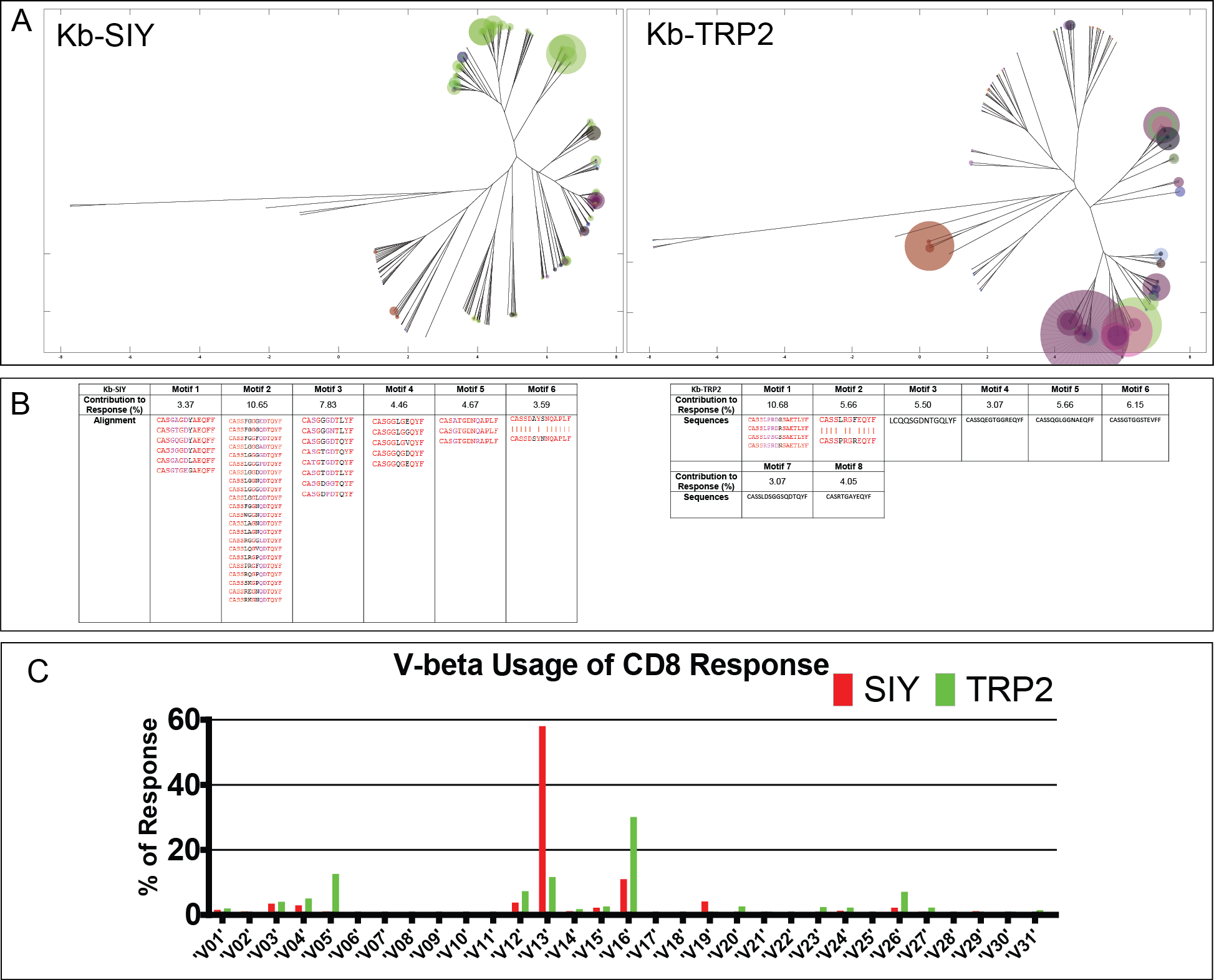
Figure 2
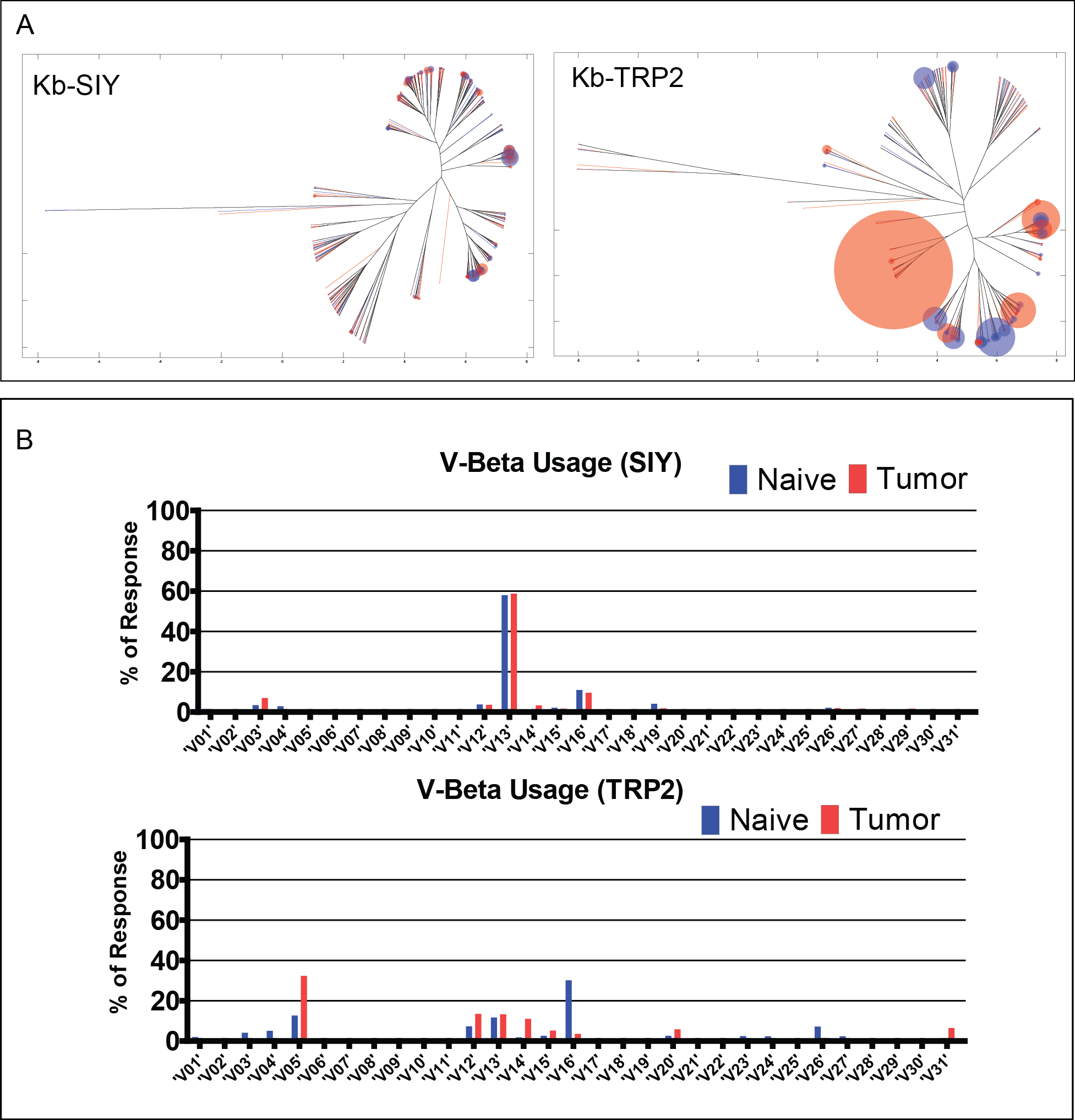
Figure 3