
Background
Before I introduce the project, you are probably wondering how I went from medical device design to cancer immunology. Over the course of my first two years of medical school, I was introduced to immunology for the first time in our pre-clinical coursework. I found the topics incredibly complex yet fascinating, and I was seeing how immunology touched every area of medicine, even orthopedics. When I had finished my second year of medical school and was about to enter the PhD years of the MD/PhD, I was faced with a critical decision about where I would take my PhD. My background in orthopedics initially pushed me to consider pursuing my doctoral work in the rapidly growing area of tissue engineering. However, my clinical experiences at the end of second year highlighted how much I enjoyed the meaningfulness of a clinical specialty such as oncology. At the time, we had been learning in medical school about the advances of immunotherapy for cancer, ushering in an era of paradigm shifting approaches in the care of cancer patients. I realized that I wanted to dedicate my life and career to the study of cancer and the care of those who had the disease. While I had no experience in cancer research, I knew that the clinical practice of oncology would provide me a lifetime of gratification and further, provide inspiration to bring to the lab each and every day. With that resolve, I took a leap of faith and joined the field of cancer immunology for my PhD.
Introduction
When I initially decided to make the jump from medical device design to cancer immunology, I faced quite a number of challenges transitioning between the two fields. It was not apparent to me at the time how I could use my previously developed skills in engineering in the field of cancer immunology. I began my work in the lab of Dr. Jonathan Schneck working for the first time on the bench in a wet lab.
As I was acclimating to this new field, I began looking for ways I could bring in my engineering skills to projects within the lab. At the time, I noticed that the field of cancer immunology was being revolutionized by technologies that generated massive amounts of data (“big data”) and there was a movement of individuals in machine learning, artificial intelligence, and statistical inference trying to understand this data. During the day I would work on projects within the lab but then at night, I would go home and spend my time teaching myself the techniques in data analysis that were being used by the field.
At the time, I remember being in lab one day when a representative from a company called Adaptive Biotechnologies came by the lab to talk to us about T-cell receptor sequencing. The representative came by my desk and asked since our lab was interrogating T-cell responses if we might be interested in this new assay they had developed. I brought this up with Dr. Schneck and the lab soon began doing TCR-Seq for projects that were ongoing in the lab. As the data began coming in, we were not exactly sure how to analyze it but I was inspired by a talk/work by Mark Davis to begin writing an algorithm to do structurally informed analysis of T-cell repertoire. The prior work by Mark Davis and Jacob Glanville was in the development of an algorithm called GLIPH that performed clustering to identify sets or “specificity groups” of TCR’s that likely recognized the same antigen.
The premise of my work was to be able to build on these concepts to visualize and quantify the diversity of a repertoire not just by the number of unique TCR’s that were present but by the structural diversity within the repertoire. Out of this, the ImmunoMap algorithm/software was born (publication). See below (after the description of the project) for my reflection on this entire transitional time in my career.
Elements of the ImmunoMap Algorithm
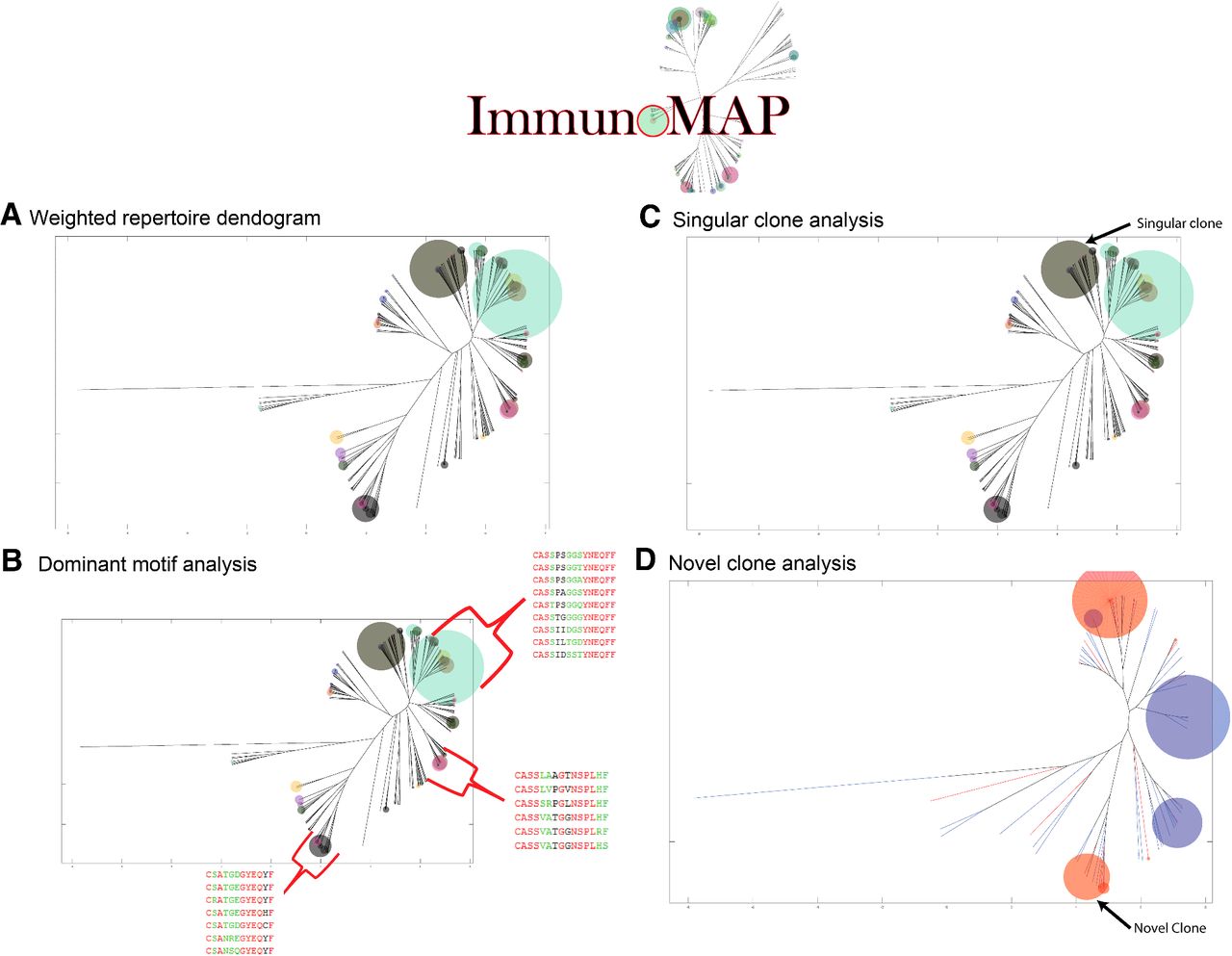
Elements of the ImmunoMap algorithm. A. Weighted repertoire dendrogram visualizes the relatedness of sequences within repertoires along with the relative frequency of CDR3 amino acid sequences. B. Dominant motif analysis clusters homologous sequences and selects for clusters contributing to significant proportion of the response. Three dominant motifs are shown that are highly represented structural motifs in this individual’s CMV response. C. Singular clone analysis defines sequences that expand significantly over the summation of all other homologous sequences. D. Novel clone analysis is implemented when comparing repertoires from different samples. A novel clone is defined as one that expands significantly over the summation of all homologous sequences in the other sample.
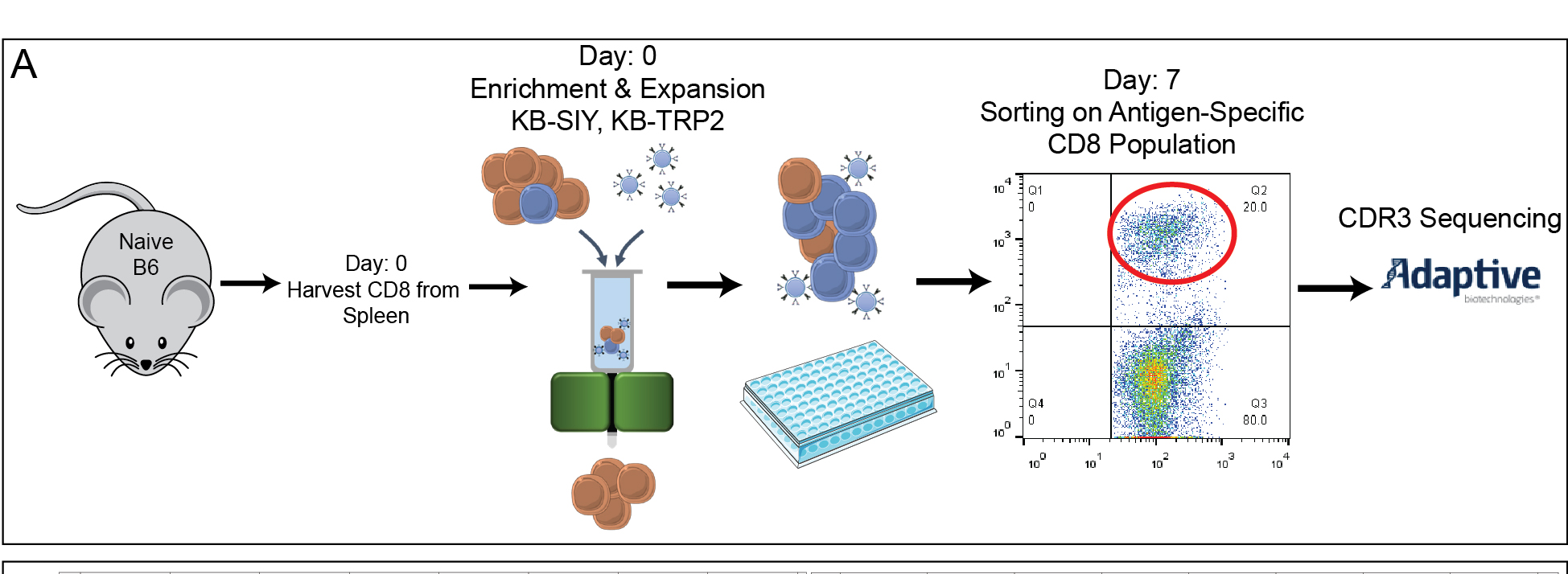
Antigen-Specific T-cell repertoires
The first question we wanted to answer in the paper was to characterize the T-cell repertoire between a self (TRP2) and non-self (SIY) murine antigen. We employed an established technique in the lab to enrich & expand antigen-specific T-cells before sequencing them.
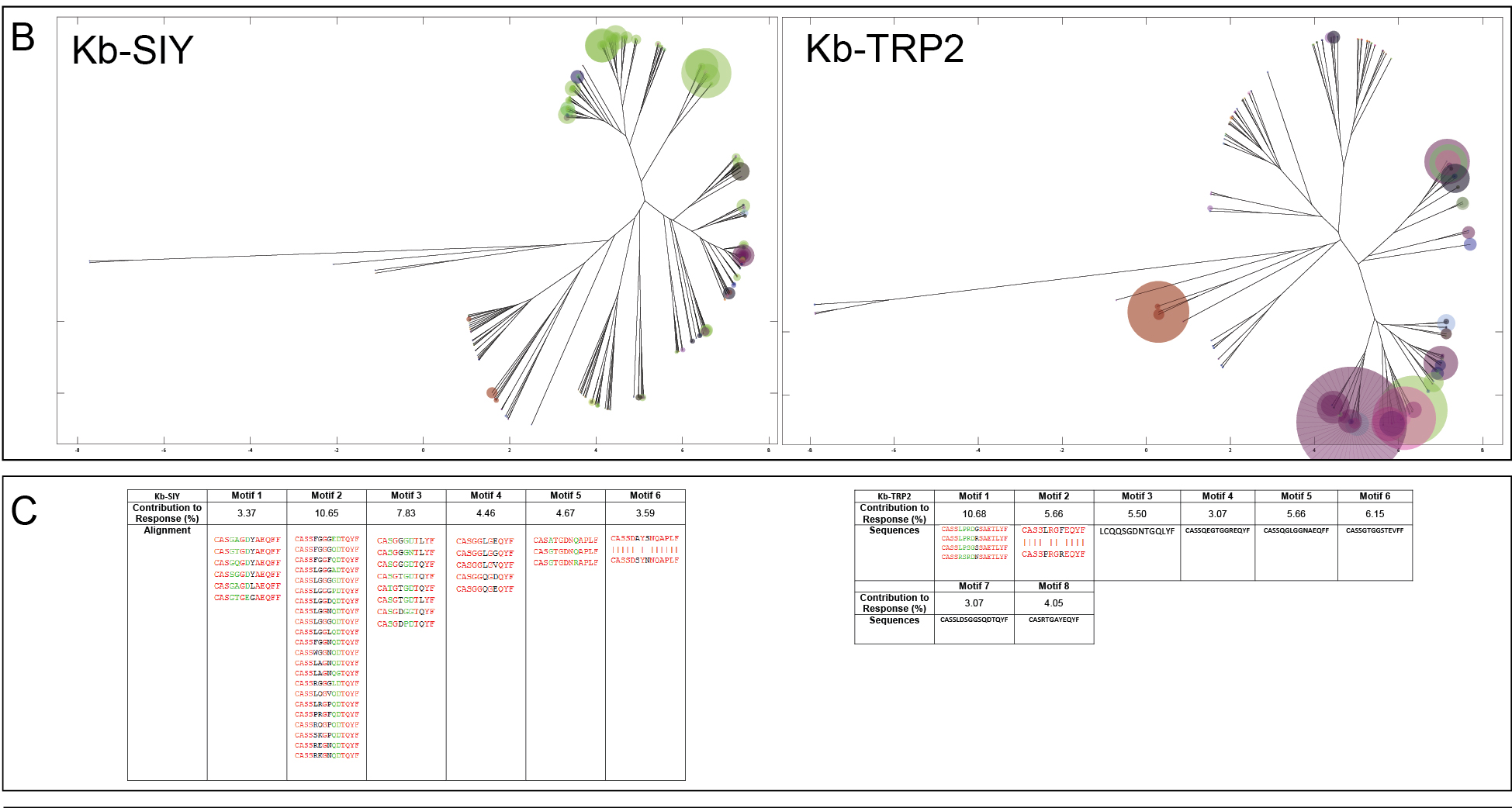
When looking at the repertoires between SIY and TRP2, we noticed that fundamentally while the non-self antigen (SIY) generated far more unique TCR’s (and thus a higher shannon’s entropy), those TCR’s were far more conserved than those that responded to TRP2. This suggested to us that this could be due to a form of selection against self-reactive TCR’s.
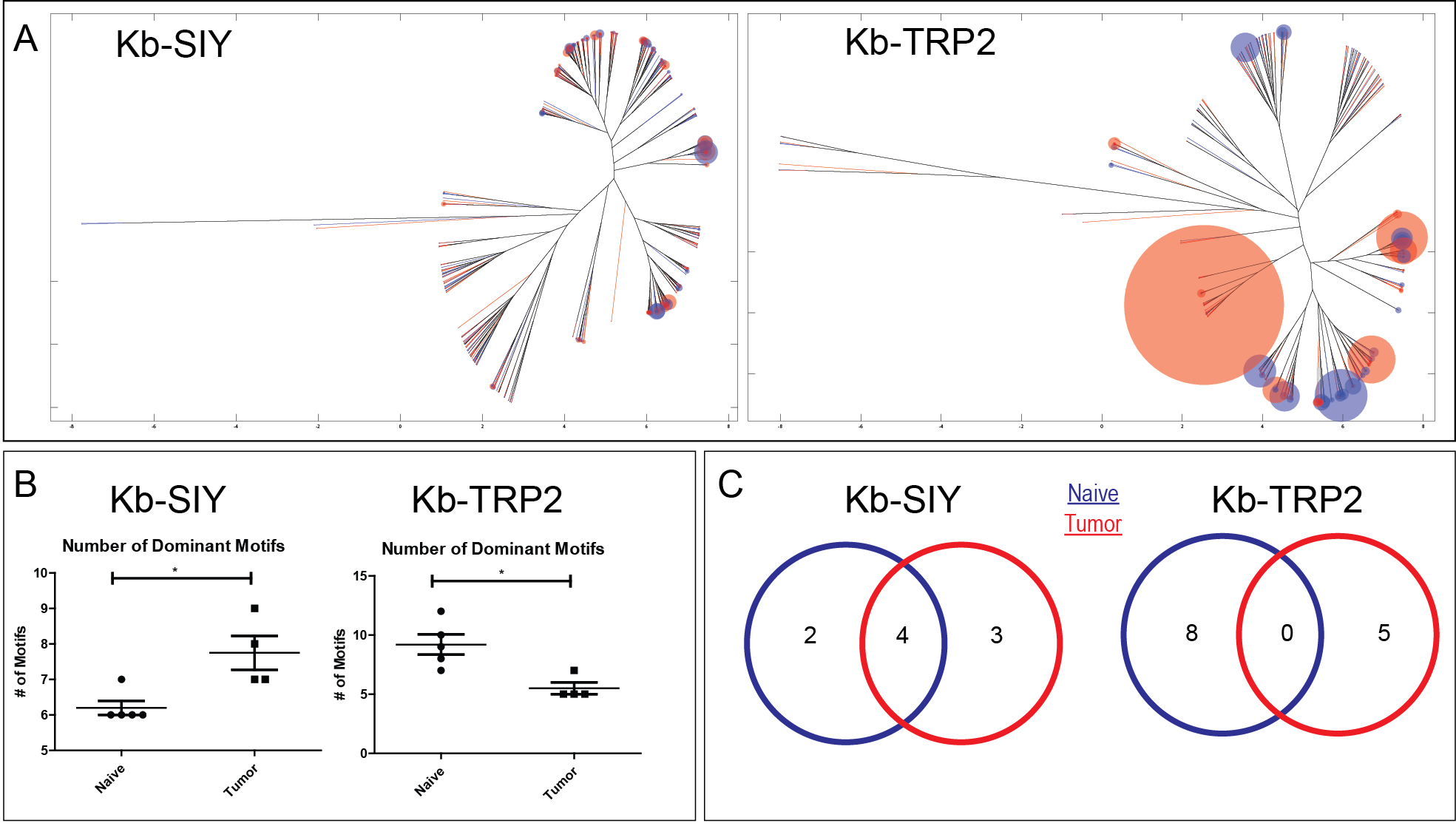
Effects of tumor on antigen-specific repertoire
As these two epitopes are often used as targets within the B6/B16 murine model, we wanted to next interrogate how the presence of tumors expressing these epitopes altered the peripheral repertoire. We noted two key findings. First, the effect of tumor seemed to prime novel structural responses to the SIY epitope while actually contracting the structural response to the TRP2 epitope (B). And second, while there was shared structural concepts between the normal and tumor repertoires for the SIY epitope, there was no shared structural concepts between the normal and tumor repertoires for the TRP2 epitope (C). These findings suggest possibly more profound immune pressure on self-antigens and thus, call into question using these as targets for immunotherapy.
Characterizing the T-cell repertoire in responders to anti-PD1 immunotherapy
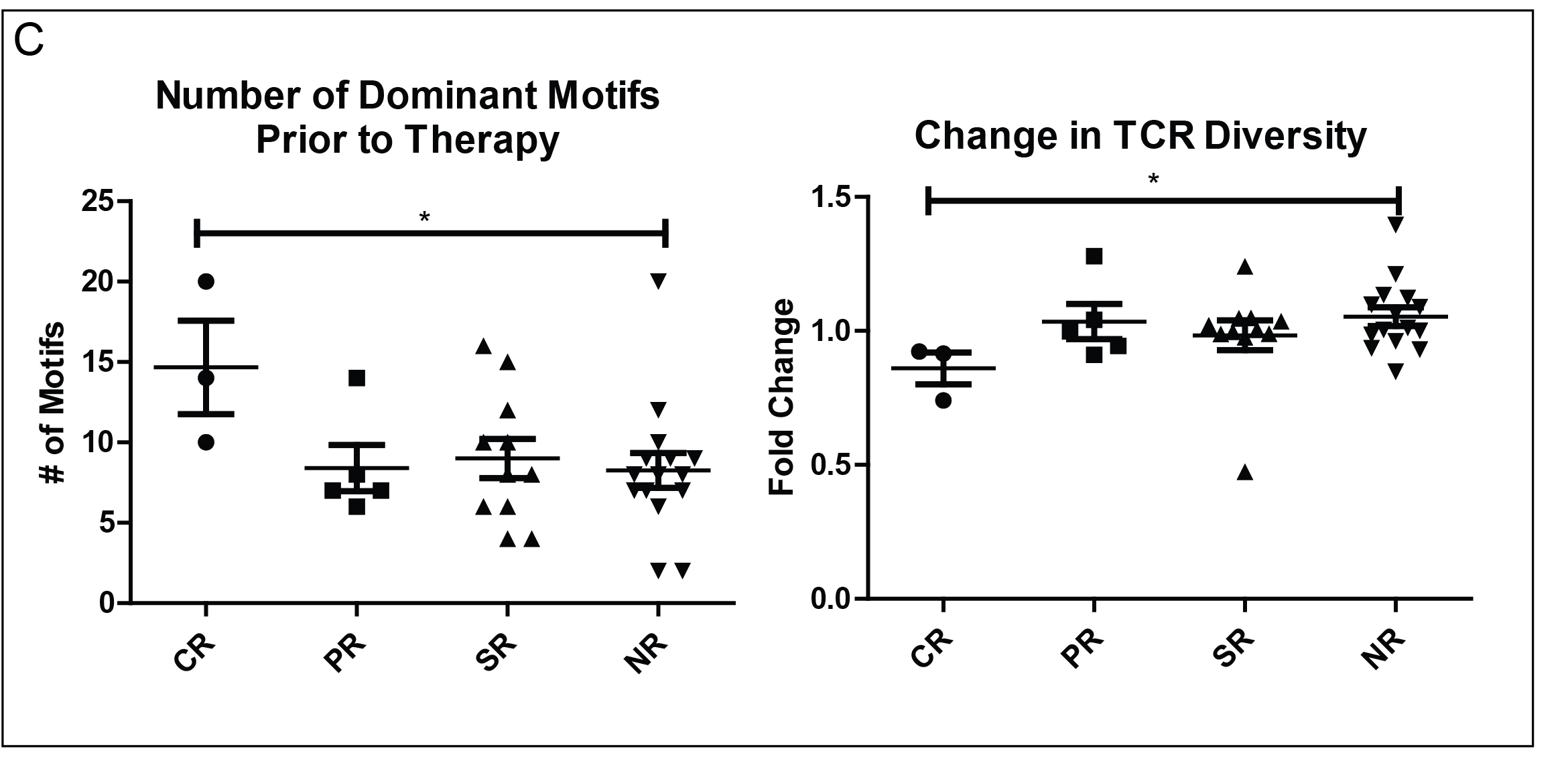
Finally, we wanted to use ImmunoMap to understand possible differences in repertoire between responders and non-responders of anti-PD1 mediated immunotherapy. We noted two important findings in our analysis. 1) Complete responders had notably more dominant motifs prior to initiation of therapy, perhaps speaking to a larger pre-existing antigenic repertoire required to generate an effective anti-tumor response. 2) Complete responders had a narrowing of their structural response after initiation of therapy suggesting that these individuals honed in on a select part of the repertoire when initiated on anti-PD1 therapy. Although our findings are significant, we note the scope of the clinical trial was limited, which impacted the distribution of clinical responses. Nevertheless, taken together, ImmunoMap analysis revealed that patients with a broader repertoire prior to therapy have a higher probability of expanding effective TCR sequences and converging on them.
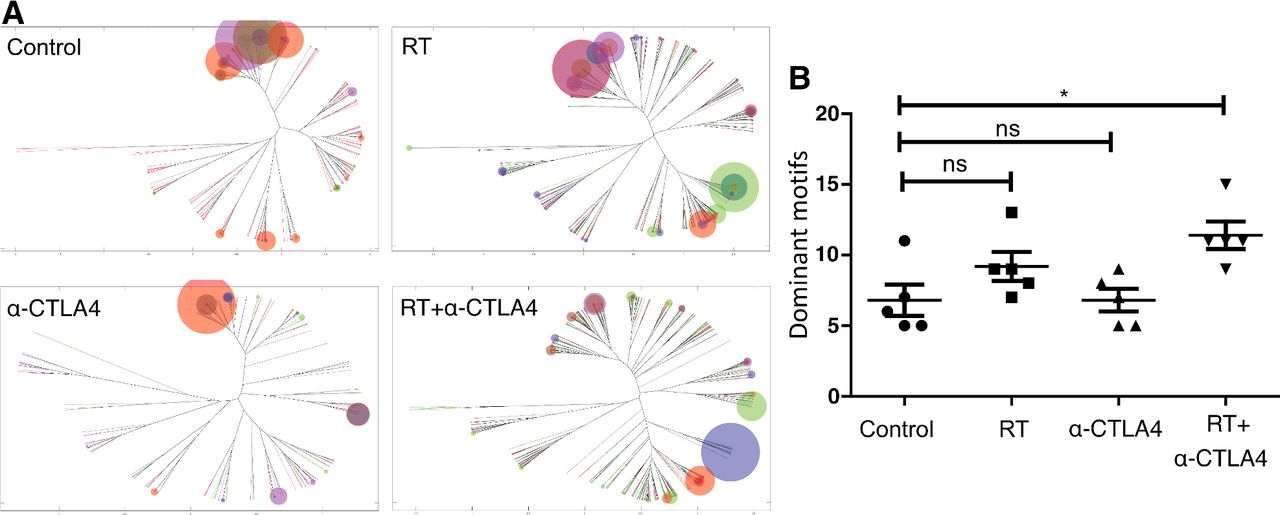
Characterizing effect of immunotherapy and radiation on T-cell repertoire in a murine model
ImmunoMap was also used in collaboration with Nils-Petter Rudqvist and Sandra Demaria at Weill Cornell Medicine where we analyzed the tumor infiltrating repertoire after various therapeutic interventions. We noted in our publication that the combination of immunotherapy and radiotherapy resulted in the largest diversification of the repertoire which was also correlated to the best response to therapy.
Reflection
When I first came into the field of cancer immunology, I would be lying if I said it was an easy transition. With a background in biomechanical engineering and mathematics, it was difficult to see any connection between engineering and cancer immunology. A few months after making the transition, I remember feeling so out of my element in the lab that I began questioning whether I had made the right decision to leave what I was familiar with for this new area of interest. And for many months, these feelings only became more present as I struggled to acclimate to the wet lab. Eventually though, the story of ImmunoMap was the story of how I was finally able to create a niche for myself where I could use my prior skills and knowledge in a way that was valuable to the field of cancer immunology. For the work of ImmunoMap, I not only wrote my first first-author paper but was invited to give oral presentations at the Society for Immunotherapy of Cancer (SITC) Annual Conference in 2016 and the Annual Meeting for the American Association for Cancer Research (AACR) in 2017. By the end of this experience, I had a clear vision of how I was going to make an impact in cancer immunology. I hoped to combine my background in engineering/mathematics (while developing further skills in machine learning and artificial intelligence) with cancer immunology in order to understand the complex interaction of the immune system and cancer in a way that would advance the diagnostics and treatment of cancer.